Enhancing a Package: Integrating Spatial Data
Follow these steps to use spatial data in a Package.
In the previous article, we used pipelines to link multiple models in our Package. In this step, we will integrate spatial data in our model and display the results using maps. Instead of having a singular linear equation representing the change over time, we will have a raster image, in which the value of each cell in the raster experiences a change over time. The value of the cell will become the new intercept in the linear equation (b in y=mt+b, where t represents time), such that each cell will experience the same slope, but will have different "starting points".
These enhancements will be part of a new Package, called helloworldSpatial, within the helloworldEnhanced suite of Packages. This Package builds off the previous Package in the suite, helloworldPipeline. Follow these steps to set up this new Package:
- Create a new subfolder within your working folder (e.g. c:\temp\helloworldSpatial)
- If creating a model in R, copy the package.xml, model.R, and model2.R files from the helloworldPipeline subfolder into the new helloworldSpatial subfolder
- If creating a model in Python, copy the package.xml, model.py, and model2.py files from the helloworldPipeline subfolder into the new helloworldSpatial subfolder.
Note
The final materials for this article can be downloaded using the following links:
Step 1 - Add Raster Inputs and Outputs to the XML Configuration File
Note
The SyncroSim XML Package has the following structure:
- XML version and Package name
- Primary Transformer and Chart Transformer
- Run Control
- First Model Inputs
- First Model Outputs / Second Model Inputs
- Second Model Ouputs
- First Transformer and Second Transformer
- Layouts (Results Transformer, Library Datafeeds, Scenario Datafeeds, Charts, Maps)
Datafeeds and Datasheets
We will need to update the input and output datafeeds so that the input datafeed can accept raster images as input and the output datafeed (in this case, the intermediate datafeed) can return raster images as output. For both the input and intermediate datasheets, add a new column with the following new attributes set to "True": isExternal and isRaster. The isExternal attribute allows the input data to be specified by a file external to the SyncroSim software. The isRaster column clarifies that this external data will be a raster image. We will call this column InterceptRasterFile. Make sure to set the dataType to "String" since the data will be a file path. In the intermediate datasheet, we also need to add an extra attribute to the InterceptRasterFile column, called bandColumn. This attribute points to another new column in the datasheet (called Band) that specifies the number of layers in the raster image. We will add this Band column as an optional column as well. Notice how we have added a new attribute to this column called allowDbNull - this attribute indicates that NULL values are allowed in the column. Below are the updated input and intermediate datafeeds in the package.xml:
<!--First Model Inputs-->
<datafeed name="InputDatafeed" displayName="Inputs" dataScope="Scenario">
<datasheets>
<datasheet name="InputDatasheet" isSingleRow="True">
<columns>
<column name="InputDatasheetID" dataType="Integer" isPrimary="True"/>
<column name="ScenarioID" dataType="Integer"/>
<column name="mMean" dataType="Double" displayName="Slope Distribution Mean"/>
<column
name="mSD"
dataType="Double"
displayName="Slope Distribution Standard Deviation"/>
<column
name="InterceptRasterFile"
displayName="Intercept Raster File"
dataType="String"
isExternalFile="True"
isRaster="True"/>
</columns>
</datasheet>
</datasheets>
</datafeed>
<!--First Model Outputs / Second Model Inputs-->
<datafeed
name="IntermediateDatafeed"
displayName="Intermediate Outputs"
dataScope="Scenario">
<datasheets>
<datasheet name="IntermediateDatasheet">
<columns>
<column
name="IntermediateDatasheetID"
dataType="Integer"
isPrimary="True"/>
<column name="ScenarioID" dataType="Integer"/>
<column name="Iteration" dataType="Integer"/>
<column name="Timestep" dataType="Integer" displayName="Timestep"/>
<column name="y" dataType="Double" displayName="Value for y"/>
<column
name="OutputRasterFile"
displayName="Output Raster File"
dataType="String"
isExternalFile="True"
isRaster="True"
bandColumn="Band"/>
<column
name="Band"
displayName="Band"
dataType="Integer"
allowDbNull="True"
isOptional="True"/>
</columns>
</datasheet>
</datasheets>
</datafeed>
Layouts
We will update the information in the <layouts> to add an option for viewing the spatial data as maps. First, we will add a Map Transformer from the corestime built-in SyncroSim Core to the Result Transformer layout. This will create a Map tab in the Results Viewer within the SyncroSim Windows Interface. See the updated Result Transformer layout below:
<!--Results Transformer Layout-->
<layout name="coreforms_ResultTransformers">
<item name="corestime_ChartTransformer"/>
<item name="corestime_MapTransformer"/>
</layout>
Lastly, we will add a Maps layout from the corestimeforms built-in SyncroSim Core. This layout stores information about which data will be used to construct the map(s). For our map, we want to specify that the data is coming from the IntermediateDatasheet and the column containing the raster output is called OutputRasterFile. Here is the new layout added to the end of the package.xml:
<!--Maps Layout-->
<layout name="corestimeforms_Maps" configurationSheet="RunControl">
<item
name="RasterMap"
displayName="Output Raster Map"
dataSheet="IntermediateDatasheet"
column="OutputRasterFile"/>
</layout>
Step 2 - Download Input Raster Data
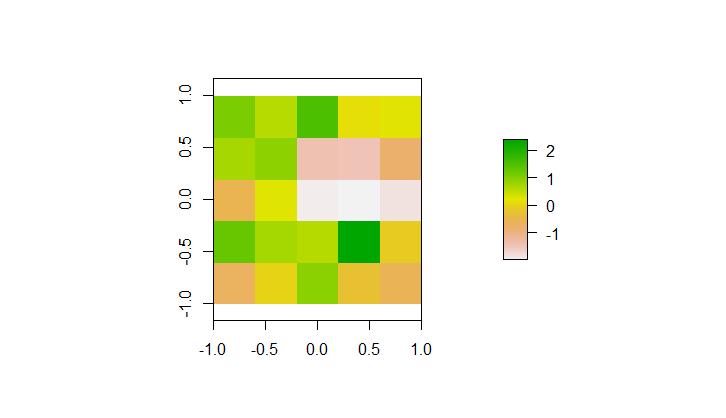
We need some simple spatial data to use in our model. Download the input-raster.tif file from the helloworldEnhanced GitHub repository and store it somewhere accessible. This image is a simple 5x5 raster, where each cell corresponds to a value taken from a normal distribution with a mean of 0 and standard deviation of 1. However, any raster image should work for this example.
Step 3 - Modify the Model Scripts
To be able to use spatial data, we will need to add some new information to the model.R script. First, we will add a line of code to load the raster library so we can read and perform calculations on raster images.
library(raster) # Load raster package
Next, retrieve the environment transfer directory for storing the raster information. The ssimEnvironment() function from rsyncrosim retrieves all environment variables for the SyncroSim session, including the transfer directory. The transfer directory is used for temporary storage, and is automatically deleted once the run has completed.
# Retrieve the transfer directory for storing output rasters
e <- ssimEnvironment()
transferDir <- e$TransferDirectory
Now we will load the input values. To load the input slope mean (mMean) and standard deviation (mSD) values we can use the same method as before. To load the input raster image, we will use the datasheetRaster() function from the rsyncrosim R package.
# Load scenario's input datasheet from SyncroSim library into R dataframe
myInputDataframe <- datasheet(myScenario,
name = "helloworldEnhanced_InputDatasheet")
# Extract model inputs from complete input dataframe
mMean <- myInputDataframe$mMean
mSD <- myInputDataframe$mSD
# Load raster input
rasterMap <- datasheetRaster(myScenario,
datasheet = "helloworldEnhanced_InputDatasheet",
column = "InterceptRasterFile")
Finally, we will add the raster calculations inside the iteration for loop. We will apply our linear equation to each cell in the input raster using the calc() function. Since timesteps is a vector, using timesteps in this function in combination with the forceapply = TRUE parameter will result in a vector of rasters with the same length as timesteps.
# Use each cell in the raster as the intercept in linear equation
newRasterMaps <- calc(rasterMap, function(b) m * timesteps + b,
forceapply = TRUE)
Next, we will get our value for y by calculating the sum of all cells in each raster. We can use the cellStats() function from the raster package to return a vector of these sums for each raster across all timesteps.
# The y value will be the sum of all the cells in each raster
y <- cellStats(newRasterMaps, stat = 'sum')
We want to store each of these new raster images in the transfer directory we retrieved earlier, so we will create file paths corresponding to the iteration and timestep of each new raster, and then write the images to these file paths.
# Add the new raster for this timestep/iteration to the output
newRasterNames <- file.path(paste0(transferDir,
"/rasterMap_iter", iter, "_ts",
timesteps, ".tif"))
writeRaster(newRasterMaps, filename = newRasterNames,
format = "GTiff", overwrite = TRUE, bylayer = TRUE)
Lastly, we will create a temporary dataframe with the Iteration, Timestep, y, and OutputRasterFile information, and add this dataframe as a new row to the output datasheet.
# Store the relevant outputs in a temporary dataframe
tempDataframe <- data.frame(Iteration = iter,
Timestep = timesteps,
y = y,
OutputRasterFile = newRasterNames)
# Copy output into this R dataframe
myOutputDataframe <- addRow(myOutputDataframe, tempDataframe)
}
The complete for loop with all updated components is shown below:
# For loop through iterations
for (iter in runSettings$MinimumIteration:runSettings$MaximumIteration) {
# Extract a slope value from normal distribution
m <- rnorm(n = 1, mean = mMean, sd = mSD)
# Use each cell in the raster as the intercept in linear equation
newRasterMaps <- calc(rasterMap, function(b) m * timesteps + b,
forceapply = TRUE)
# The y value will be the sum of all the cells in each raster
y <- cellStats(newRasterMaps, stat = 'sum')
# Add the new raster for this timestep/iteration to the output
newRasterNames <- file.path(paste0(transferDir,
"/rasterMap_iter", iter, "_ts",
timesteps, ".tif"))
writeRaster(newRasterMaps, filename = newRasterNames,
format = "GTiff", overwrite = TRUE, bylayer = TRUE)
# Store the relevant outputs in a temporary dataframe
tempDataframe <- data.frame(Iteration = iter,
Timestep = timesteps,
y = y,
OutputRasterFile = newRasterNames)
# Copy output into this R dataframe
myOutputDataframe <- addRow(myOutputDataframe, tempDataframe)
}
The model2.R script will remain the same.
Step 4 - Rebuild the Package
Use the Package Manager to rebuild the Package using the new package.xml and model.R or model.py files shown above (see Step 4 in the Building a Package article).
Step 5 - Set Model Inputs
- Start SyncroSim.
- Create a new library based on the helloworldEnhanced Package.
- Within the Library Properties, set the location of the R or Python executable in the R Configuration or Python Configuration tab.
- Create a new Scenario - Right-click in the Library Explorer and select New > Scenario from the context menu. Rename this scenario to Spatial.
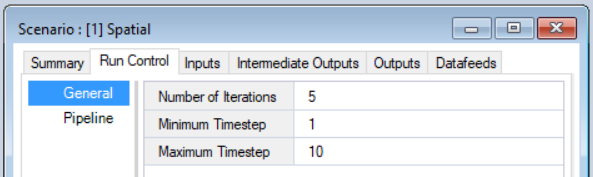
- Edit the Scenario Run Control - Within the Spatial Scenario, navigate to the Run Control - General tab. Set the Number of Iterations, Minimum Timestep, and Maximum Timestep values for your model. Make sure to also set the order of Transformers in the Pipeline tab.
- Edit the Scenario Pipeline - Now, on the Run Control - Pipeline tab (see previous step), add the Transformers to the Stage section and specify the Run Order.
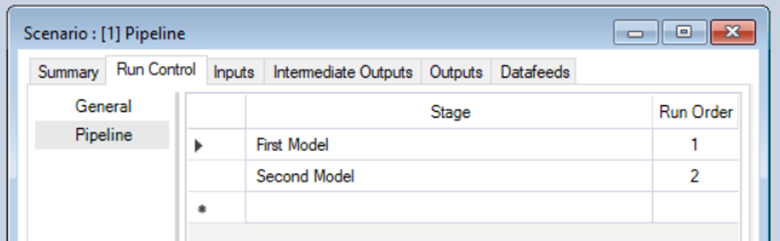
- Edit the Scenario Inputs - Choose values for mMean and mSD. Click on the folder beside Intercept Raster File and select the "input-raster.tif" we saved earlier.
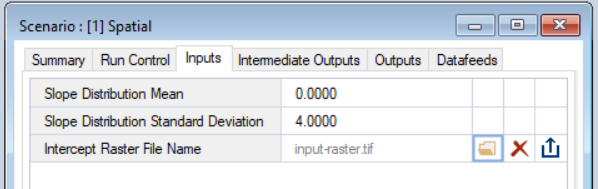
- Save the Scenario inputs - Save changes made to the Scenario inputs.
Step 6 - Run the Model
Right-click on this Spatial Scenario in the Library Explorer and select Run to run this Scenario.
Step 7 - View the Results
- Once the run is complete, return to the Library Explorer. Expand the node beside the Spatial scenario to reveal a Results folder containing your results, then expand the node beside the Results folder to show the newly generated date/time stamped Results Scenario.
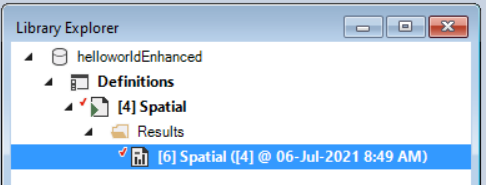
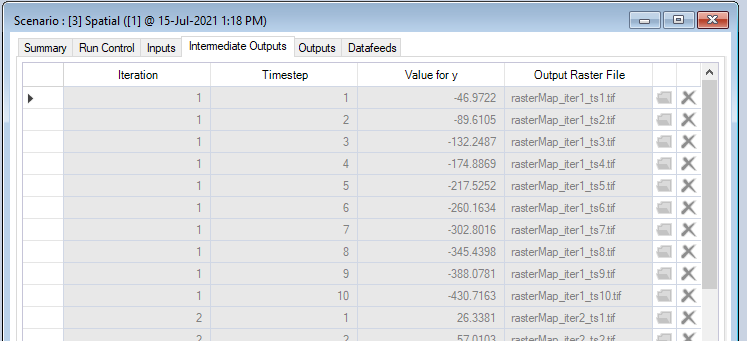
- Right-click on this Results Scenario and select Properties to view the details of this Results Scenario; you will find the raster files that correspond to each iteration and timestep under the Intermediate Outputs tab. There is also a column for the Value for y, displaying the total sum of cells for each raster for each iteration and timestep.
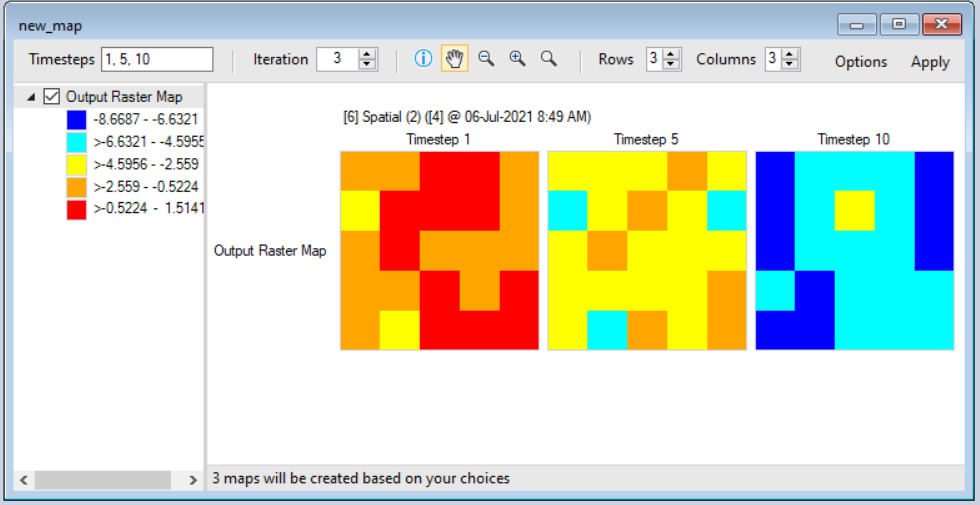
- To view the resulting raster images as maps, click on the Maps tab in the Results Viewer, and click Create a new map. Give your map a name.
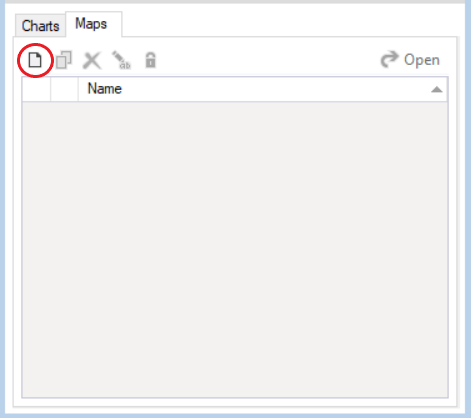
- In the Create a new map pane, select Output Raster Map from the left-hand side. Click Apply to view the map of the last timestep for the first iteration. You can view the maps at the 1st, 5th, and 10th timesteps across the simulation by setting the Timesteps value in the top toolbar to "1,5,10". You can also change the iteration by clicking on the up or down arrows next to Iteration in the top toolbar. After making these changes, click Apply again to view the updated maps. To make further modifications to your maps, such as creating a custom colour legend, see the Customizing a Map tutorial.